SimulConsult puts expert knowledge at your fingertips
Benefits of getting the differential diagnosis right the first time
Improves accuracy
In NIH-funded studies, physician use of the software reduces diagnostic errors by up to 75%.
Makes you feel wise
With the wisdom of the community from many books, articles and other resources at your fingertips, you can diagnose confidently even beyond your specialty.
Saves you time
It takes 7 minutes on average to use the software, but saves 30 minutes or more through automated documentation and time no longer spent searching through narrative articles.
Unique database
The human curated database has 9,700 diagnoses, and is most complete for genetics, neurology and increasingly pediatric rheumatology. It includes the genes with convincing human disease phenotypes in OMIM and the diseases in GeneReviews.
Robust model
The computational software takes into account pertinent negatives, temporal information, and cost of tests — ignored in other diagnostic approaches. It transforms medical diagnosis by lowering costs, reducing errors, improving documentation and eliminating the medical diagnostic odysseys. See the Frequently Asked Questions for particular capabilities you may need.
Explainable AI approach
SimulConsult uses an AI-based statistical pattern-matching approach that takes into account the age of onset and offset of the findings in each disease. The software is distinctive in revealing its logic in a series of Assess and Profile tools, so that the logic is “explainable”, a capability considered by clinicians as equally or more important than the ability to identify consistently the correct diagnosis.
Intuitive user interface
Designed for doctors by doctors, the user interface mimics how great diagnosticians think and how all of us were trained to do diagnosis, so getting up to speed is quick and easy.
Mobile
The web site and software work from mobile devices as well as personal computers. There is no app to download. Just access the diagnostic tool from the SimulConsult web site. Your license lets you use the tool from your mobile device as well as your computer. It works on larger phones, but a small tablet offers the best mix of mobility and convenience.
Thousands of links
Findings (signs, symptoms, lab results) and diseases have “Tip” hyperlinks to review articles, papers and images to help with diagnosis. Even if you have never seen a particular finding before, it may be suggested as a useful finding in “Add findings” or “Add tests” based on its ability to change the differential diagnosis. The Tip for the finding can include just-in-time information such as images or articles, or even a percentile calculator already filled in with your patient’s age and sex.
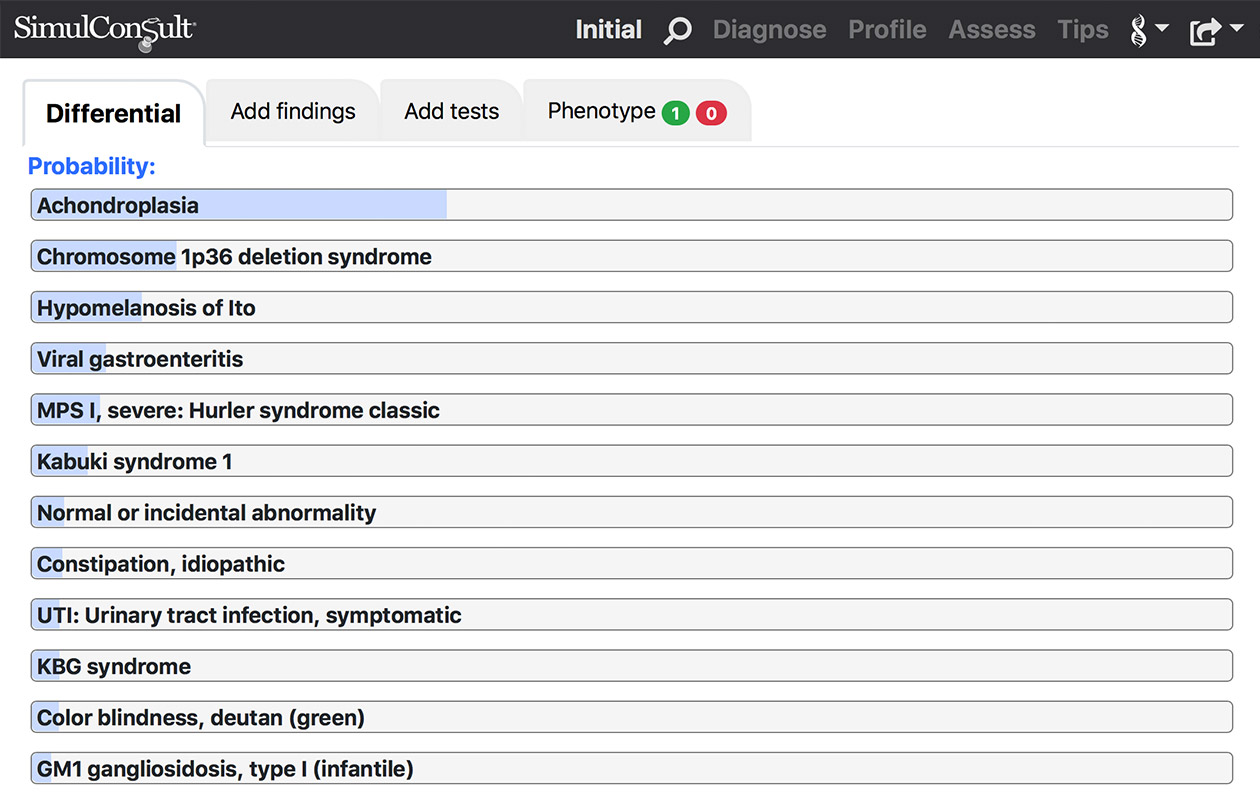
Phenome Analyzer
All the tools you need for regular diagnosis.- Differential diagnosis display
- Useful finding displays (clinical & tests)
- Add findings (positives with onset, and negatives)
- Assess diagnosis (“explainable AI”)
- Tip links about current disease and finding
- Family history: specify in a few clicks
- Patient Summary: save and reload
- Prognosis Table output explaining what to expect
- Regions of homozygosity analysis (microarrays)
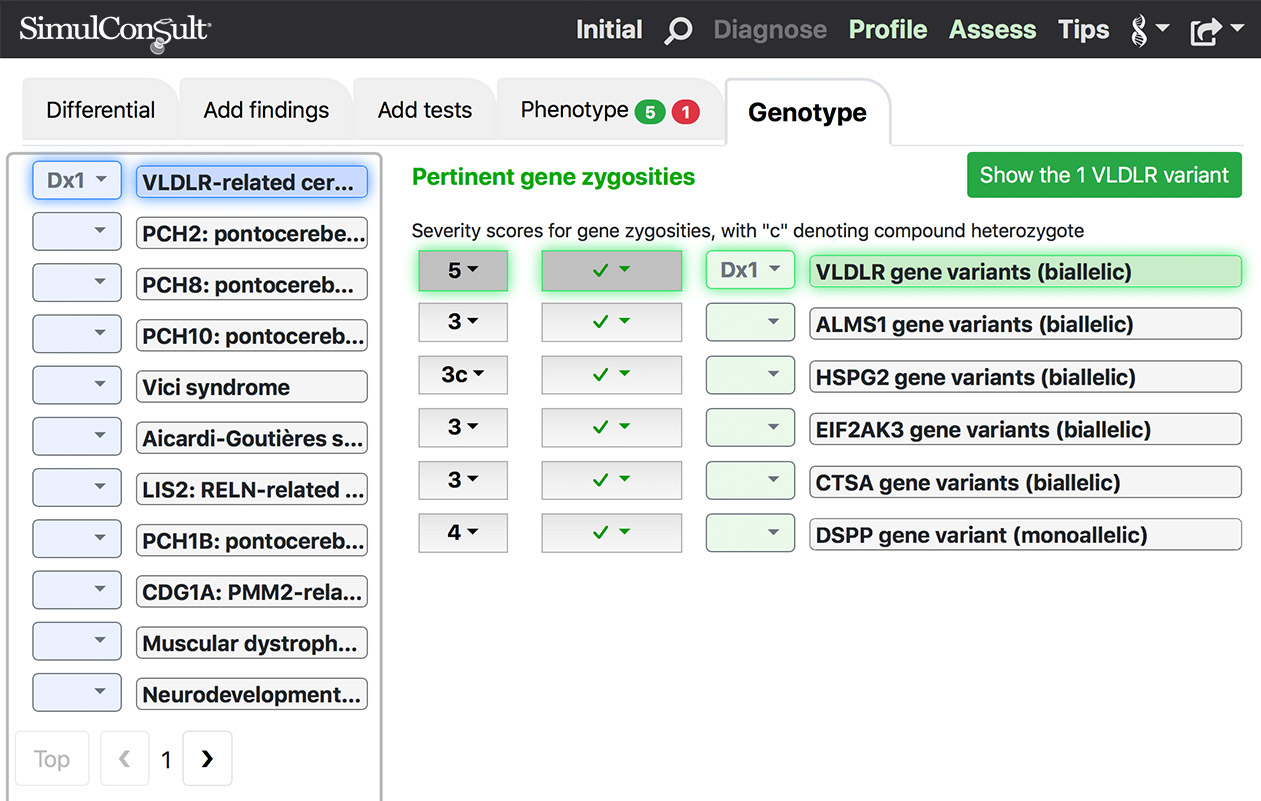
Genome-Phenome Analyzer
Adds the ability to import annotated variant tables and analyze the full genomic results.- All the features in the Phenome analyzer
- Variant table upload for proband, trio or beyond the trio
- PED file upload for beyond the trio
- Picks out the gene zygosities most pertinent for your patient
- Genome report output including up to 4 diagnoses
- Discovery report for genes with unknown clinical phenotype
Start Patient
Specify or import:
- Age
- Sex
- Care setting (primary, specialist, tertiary)
- Emergence speed
- Family history in a few clicks
Add Findings
Phenome Analyzer
Findings can include not only presence, but also onset or absence
Add findings by search
Add findings using saving specialty-based core finding lists
Add findings from useful finding list (clinical or tests)
Import microarray data
Explore Differential Diagnosis
Phenome Analyzer
View evolving differential diagnosis with probabilities displayed as a bar chart
View contextually-relevant Tips (Includes OMIM, Orphanet, Gene Reviews, Elements of Morphology, images, videos, etc.)
Assess & Plan
Phenome Analyzer
Assess fit of selected disease to patient findings
See a profile of the disease by age of patient
View profile of a finding across diseases generally or those in the differential diagnosis
Identify most useful tests to order
Save/Reopen Document
Phenome Analyzer
Save Patient Summary: human readable PDF, but machine loadable
Save Chart Note Template (History, Exam, Assessment, Plan)
Prognosis Table to explain natural history of the diagnosis to patient and referring physician
